2015
CIBC.
Note: Data Sets: NCRR Center for Integrative Biomedical Computing (CIBC) data set archive. Download from: http://www.sci.utah.edu/cibc/software.html, 2015.
CIBC.
Note: Cleaver: A MultiMaterial Tetrahedral Meshing Library and Application. Scientific Computing and Imaging Institute (SCI), Download from: http://www.sci.utah.edu/cibc/software.html, 2015.
Y. Gao, L. Zhu, J. Cates, R. S. MacLeod, S. Bouix,, A. Tannenbaum.
“A Kalman Filtering Perspective for Multiatlas Segmentation,” In SIAM J. Imaging Sciences, Vol. 8, No. 2, pp. 1007-1029. 2015.
DOI: 10.1137/130933423
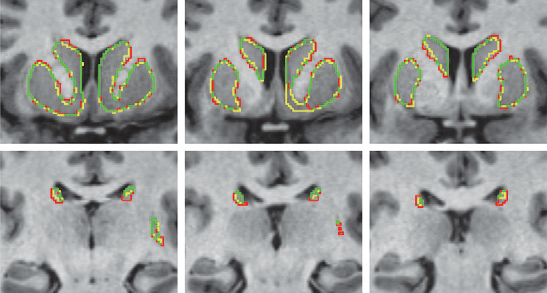
K. Gillette, J.D. Tate, B. Kindall, P. Van Dam, E. Kholmovski, R.S. MacLeod.
“Generation of Combined-Modality Tetrahedral Meshes,” In Computing in Cardiology, 2015.
Registering and combining anatomical components from different image modalities, like MRI and CT that have different tissue contrast, could result in patient-specific models that more closely represent underlying anatomical structures.
In this study, we combined a pair of CT and MRI scans of a pig thorax to make a tetrahedral mesh and compared different registration techniques including rigid, affine, thin plate spline morphing (TPSM), and iterative closest point (ICP), to superimpose the segmented bones from the CT scan on the soft tissues segmented from the MRI. The TPSM and affine-registered bones remained close to, but not overlapping, important soft tissue.
Simulation models, including an ECG forward model and a defibrillation model, were computed on generated multi-modality meshes after TPSM and affine registration and compared to those based on the original torso mesh.
CIBC.
Note: ImageVis3D: An interactive visualization software system for large-scale volume data. Scientific Computing and Imaging Institute (SCI), Download from: http://www.imagevis3d.org, 2015.
C.R. Johnson.
“Computational Methods and Software for Bioelectric Field Problems,” In Biomedical Engineering Handbook, 4, Vol. 1, Ch. 43, Edited by J.D. Bronzino and D.R. Peterson, CRC Press, pp. 1--28. 2015.
Computer modeling and simulation continue to become more important in the field of bioengineering. The reasons for this growing importance are manyfold. First, mathematical modeling has been shown to be a substantial tool for the investigation of complex biophysical phenomena. Second, since the level of complexity one can model parallels the existing hardware configurations, advances in computer architecture have made it feasible to apply the computational paradigm to complex biophysical systems. Hence, while biological complexity continues to outstrip the capabilities of even the largest computational systems, the computational methodology has taken hold in bioengineering and has been used successfully to suggest physiologically and clinically important scenarios and results.
This chapter provides an overview of numerical techniques that can be applied to a class of bioelectric field problems. Bioelectric field problems are found in a wide variety of biomedical applications, which range from single cells, to organs, up to models that incorporate partial to full human structures. We describe some general modeling techniques that will be applicable, in part, to all the aforementioned applications. We focus our study on a class of bioelectric volume conductor problems that arise in electrocardiography (ECG) and electroencephalography (EEG).
We begin by stating the mathematical formulation for a bioelectric volume conductor, continue by describing the model construction process, and follow with sections on numerical solutions and computational considerations. We continue with a section on error analysis coupled with a brief introduction to adaptive methods. We conclude with a section on software.
C.R. Johnson.
“Visualization,” In Encyclopedia of Applied and Computational Mathematics, Edited by Björn Engquist, Springer, pp. 1537-1546. 2015.
ISBN: 978-3-540-70528-4
DOI: 10.1007/978-3-540-70529-1_368
CIBC.
Note: map3d: Interactive scientific visualization tool for bioengineering data. Scientific Computing and Imaging Institute (SCI), Download from: http://www.sci.utah.edu/cibc/software.html, 2015.
K.S. McDowell, S. Zahid, F. Vadakkumpadan, J.J. Blauer, R.S. MacLeod, N.A. Trayanova.
“Virtual Electrophysiological Study of Atrial Fibrillation in Fibrotic Remodeling,” In PLoS ONE, Vol. 10, No. 2, pp. e0117110. February, 2015.
DOI: 10.1371/journal.pone.0117110
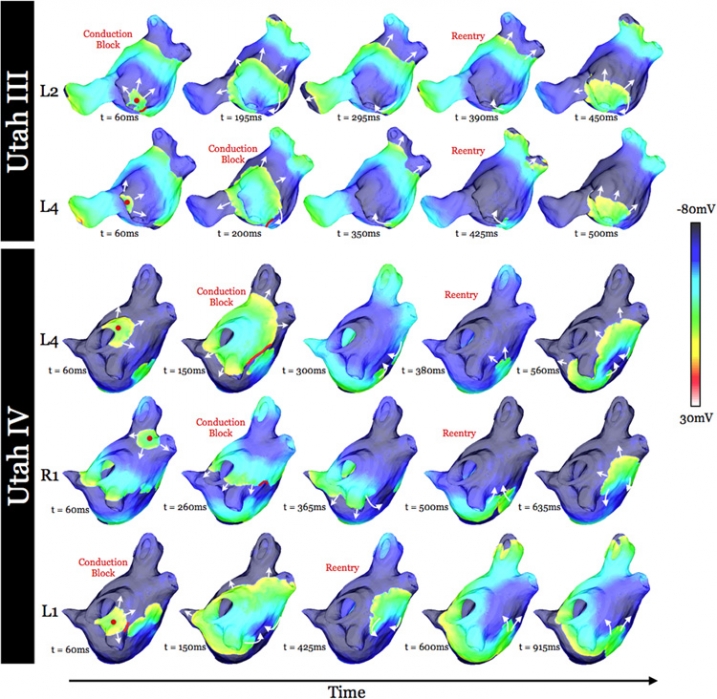
K. S. McDowell, S. Zahid, F. Vadakkumpadan, J. Blauer, R. S. MacLeod, N. A. Trayanova.
“Virtual Electrophysiological Study of Atrial Fibrillation in Fibrotic Remodeling,” In PLoS ONE, Vol. 10, No. 2, Public Library of Science, pp. 1-16. May, 2015.
DOI: doi.org/10.1371/journal.pone.0117110
Research has indicated that atrial fibrillation (AF) ablation failure is related to the presence of atrial fibrosis. However it remains unclear whether this information can be successfully used in predicting the optimal ablation targets for AF termination. We aimed to provide a proof-of-concept that patient-specific virtual electrophysiological study that combines i) atrial structure and fibrosis distribution from clinical MRI and ii) modeling of atrial electrophysiology, could be used to predict: (1) how fibrosis distribution determines the locations from which paced beats degrade into AF; (2) the dynamic behavior of persistent AF rotors; and (3) the optimal ablation targets in each patient. Four MRI-based patient-specific models of fibrotic left atria were generated, ranging in fibrosis amount. Virtual electrophysiological studies were performed in these models, and where AF was inducible, the dynamics of AF were used to determine the ablation locations that render AF non-inducible. In 2 of the 4 models patient-specific models AF was induced; in these models the distance between a given pacing location and the closest fibrotic region determined whether AF was inducible from that particular location, with only the mid-range distances resulting in arrhythmia. Phase singularities of persistent rotors were found to move within restricted regions of tissue, which were independent of the pacing location from which AF was induced. Electrophysiological sensitivity analysis demonstrated that these regions changed little with variations in electrophysiological parameters. Patient-specific distribution of fibrosis was thus found to be a critical component of AF initiation and maintenance. When the restricted regions encompassing the meander of the persistent phase singularities were modeled as ablation lesions, AF could no longer be induced. The study demonstrates that a patient-specific modeling approach to identify non-invasively AF ablation targets prior to the clinical procedure is feasible.
I. OguzI, J. Cates, M. Datar, B. Paniagua, T. Fletcher, C. Vachet, M. Styner, R. Whitaker.
“Entropy-based particle correspondence for shape populations,” In International Journal of Computer Assisted Radiology and Surgery, Springer, pp. 1-12. December, 2015.
Purpose
Statistical shape analysis of anatomical structures plays an important role in many medical image analysis applications such as understanding the structural changes in anatomy in various stages of growth or disease. Establishing accurate correspondence across object populations is essential for such statistical shape analysis studies.
Methods
In this paper, we present an entropy-based correspondence framework for computing point-based correspondence among populations of surfaces in a groupwise manner. This robust framework is parameterization-free and computationally efficient. We review the core principles of this method as well as various extensions to deal effectively with surfaces of complex geometry and application-driven correspondence metrics.
Results
We apply our method to synthetic and biological datasets to illustrate the concepts proposed and compare the performance of our framework to existing techniques.
Conclusions
Through the numerous extensions and variations presented here, we create a very flexible framework that can effectively handle objects of various topologies, multi-object complexes, open surfaces, and objects of complex geometry such as high-curvature regions or extremely thin features.
B.R. Parmar, T.R. Jarrett, E.G. Kholmovski, N. Hu, D. Parker, R.S. MacLeod, N.F. Marrouche, R. Ranjan.
“Poor scar formation after ablation is associated with atrial fibrillation recurrence,” In Journal of Interventional Cardiac Electrophysiology, Vol. 44, No. 3, pp. 247-256. December, 2015.
Purpose
Patients routinely undergo ablation for atrial fibrillation (AF) but the recurrence rate remains high. We explored in this study whether poor scar formation as seen on late-gadolinium enhancement magnetic resonance imaging (LGE-MRI) correlates with AF recurrence following ablation.
Methods
We retrospectively identified 94 consecutive patients who underwent their initial ablation for AF at our institution and had pre-procedural magnetic resonance angiography (MRA) merged with left atrial (LA) anatomy in an electroanatomic mapping (EAM) system, ablated areas marked intraprocedurally in EAM, 3-month post-ablation LGE-MRI for assessment of scar, and minimum of 3-months of clinical follow-up. Ablated area was quantified retrospectively in EAM and scarred area was quantified in the 3-month post-ablation LGE-MRI.
Results
With the mean follow-up of 336 days, 26 out of 94 patients had AF recurrence. Age, hypertension, and heart failure were not associated with AF recurrence, but LA size and difference between EAM ablated area and LGE-MRI scar area was associated with higher AF recurrence. For each percent higher difference between EAM ablated area and LGE-MRI scar area, there was a 7–9 % higher AF recurrence (p values 0.001–0.003) depending on the multivariate analysis.
Conclusions
In AF ablation, poor scar formation as seen on LGE-MRI was associated with AF recurrence. Improved mapping and ablation techniques are necessary to achieve the desired LA scar and reduce AF recurrence.
SCI Institute.
Note: SCIRun: A Scientific Computing Problem Solving Environment, Scientific Computing and Imaging Institute (SCI), Download from: http://www.scirun.org, 2015.
CIBC.
Note: Seg3D: Volumetric Image Segmentation and Visualization. Scientific Computing and Imaging Institute (SCI), Download from: http://www.seg3d.org, 2015.
2014
J.J.E. Blauer, D. Swenson, K. Higuchi, G. Plank, R. Ranjan, N. Marrouche,, R.S. MacLeod.
“Sensitivity and Specificity of Substrate Mapping: An In Silico Framework for the Evaluation of Electroanatomical Substrate Mapping Strategies,” In Journal of Cardiovascular Electrophysiology, In Journal of Cardiovascular Electrophysiology, Vol. 25, No. 7, Note: Featured on journal cover., pp. 774--780. May, 2014.
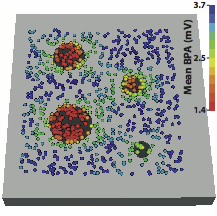
Keywords: arrhythmia, computer-based model, electroanatomical mapping, voltage mapping, bipolar electrogram
J. Bronson, J.A. Levine, R.T. Whitaker.
“Lattice cleaving: a multimaterial tetrahedral meshing algorithm with guarantees,” In IEEE Transactions on Visualization and Computer Graphics (TVCG), pp. 223--237. 2014.
DOI: 10.1109/TVCG.2013.115
PubMed ID: 24356365
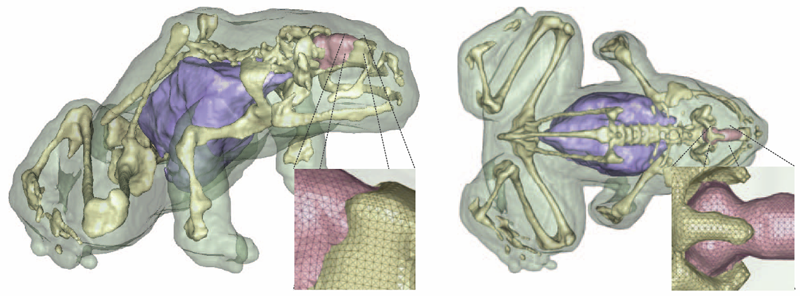
S. Eichelbaum, M. Dannhauer, M. Hlawitschka , D.H. Brooks, T.R. Knosche, G. Scheuermanna.
“Visualizing Simulated Electrical Fields from Electroencephalography and Transcranial Electric Brain Stimulation: A Comparative Evaluation,” In Neuroimage, 2014.
DOI: 10.1016/j.neuroimage.2014.04.085
Electrical activity of neuronal populations is a crucial aspect of brain activity. This activity is not measured directly but recorded as electrical potential changes using head surface electrodes (electroencephalogram - EEG). Head surface electrodes can also be deployed to inject electrical currents in order to modulate brain activity (transcranial electric stimulation techniques) for therapeutic and neuroscientific purposes. In electroencephalography and noninvasive electric brain stimulation, electrical fields mediate between electrical signal sources and regions of interest (ROI). These fields can be very complicated in structure, and are influenced in a complex way by the conductivity profile of the human head. Visualization techniques play a central role to grasp the nature of those fields because such techniques allow for an effective conveyance of complex data and enable quick qualitative and quantitative assessments. The examination of volume conduction effects of particular head model parameterizations (e.g., skull thickness and layering), of brain anomalies (e.g., holes in the skull, tumors), location and extent of active brain areas (e.g., high concentrations of current densities) and around current injecting electrodes can be investigated using visualization. Here, we evaluate a number of widely used visualization techniques, based on either the potential distribution or on the current-flow. In particular, we focus on the extractability of quantitative and qualitative information from the obtained images, their effective integration of anatomical context information, and their interaction. We present illustrative examples from clinically and neuroscientifically relevant cases and discuss the pros and cons of the various visualization techniques.
Keywords: Visualization, Bioelectric Field, EEG, tDCS, Human Brain
B. Erem P.M. van Dam, D.H. Brooks.
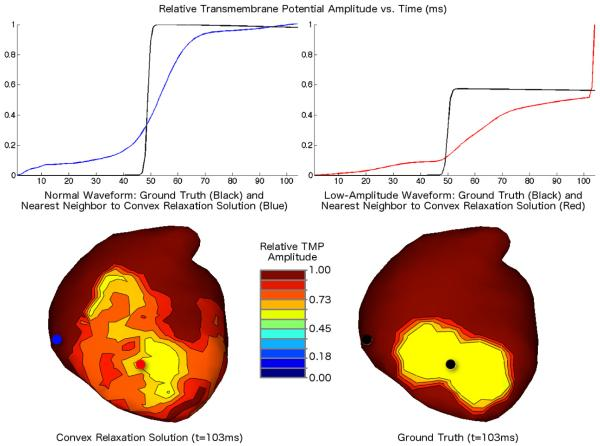
“Identifying model inaccuracies and solution uncertainties in noninvasive activation-based imaging of cardiac excitation using convex relaxation,” In IEEE Trans Med Imaging, Vol. 33, No. 4, pp. 902--912. 2014.
DOI: 10.1109/TMI.2014.2297952
PubMed ID: 24710159
PubMed Central ID: PMC3982205
B. Erem, J. Coll-Font, R.M. Orellana, P. Stovicek, D.H. Brooks.
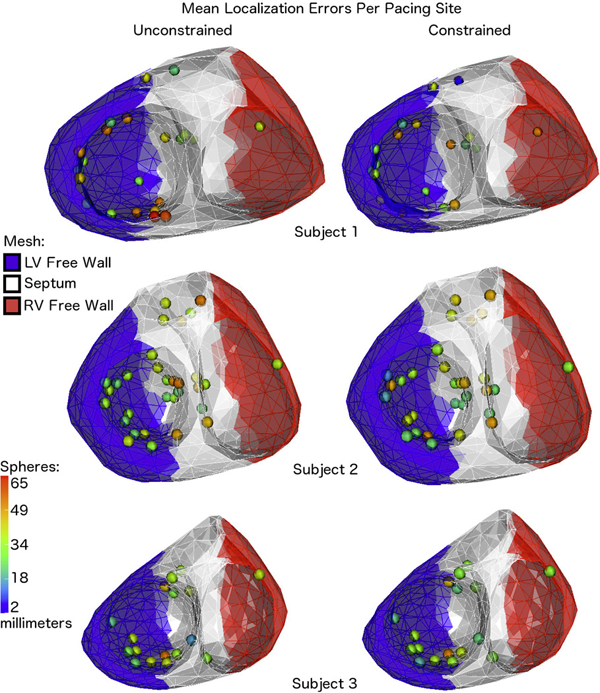
“Using transmural regularization and dynamic modeling for noninvasive cardiac potential imaging of endocardial pacing with imprecise thoracic geometry,” In IEEE Trans Med Imaging, Vol. 33, No. 3, pp. 726--738. 2014.
DOI: 10.1109/TMI.2013.2295220
PubMed ID: 24595345
PubMed Central ID: PMC3950945
T. Fogal, F. Proch, A. Schiewe, O. Hasemann, A. Kempf, J. Krüger.
“Freeprocessing: Transparent in situ visualization via data interception,” In Proceedings of the 14th Eurographics Conference on Parallel Graphics and Visualization, EGPGV, Eurographics Association, 2014.
In situ visualization has become a popular method for avoiding the slowest component of many visualization pipelines: reading data from disk. Most previous in situ work has focused on achieving visualization scalability on par with simulation codes, or on the data movement concerns that become prevalent at extreme scales. In this work, we consider in situ analysis with respect to ease of use and programmability. We describe an abstraction that opens up new applications for in situ visualization, and demonstrate that this abstraction and an expanded set of use cases can be realized without a performance cost.
Page 6 of 24